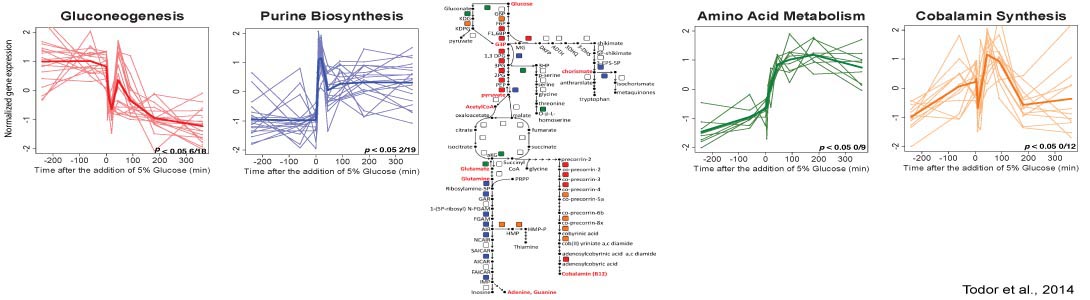
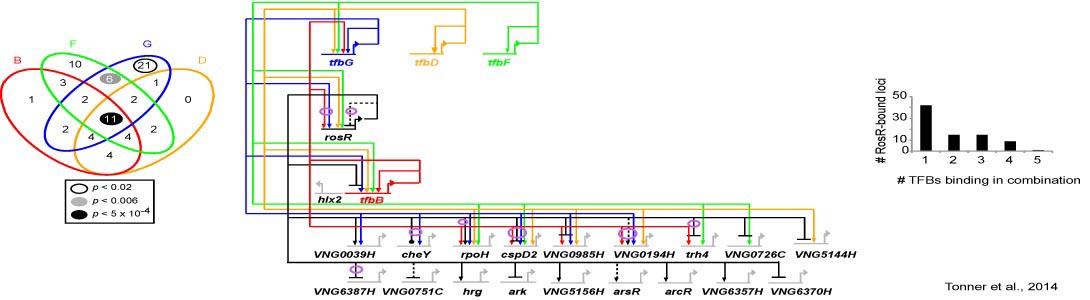
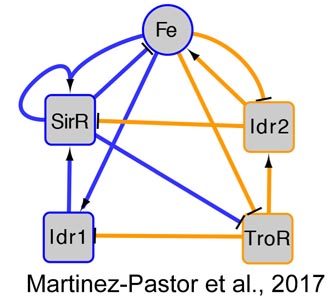
How do archaea regulate responses to extreme stress?
How do these responses evolve?
Objectives
Extremophiles thrive in deep-sea hydrothermal vents under high pressure and temperature, saturated salt lakes, and polar icecaps. Many of these organisms are members of the third domain of life, the archaea. Although archaea contribute substantially to global carbon and energy cycles, they remain understudied because they are difficult to culture and manipulate genetically. How do these microorganisms cope with an extreme and changing environment? How do they alter their genetic programs and metabolic pathways to adapt and survive changes in their unique habitats?
Central Questions
To address these questions, research in our group uses salt-adapted archaea to understand how these organisms grow in energy limited salt lakes while surviving cellular damage from heavy solar irradiation. Specifically, we use a systems biology approach to ask:
- How do transcription factors work together dynamically to regulate these responses?
- What are the molecular function and mechanisms of transcription factors in archaea?
Photography courtesy of Jim Gilbert